Plots of the Output of a Bootstrap Simulation
plot.boot.RdThis takes a bootstrap object and produces plots for the bootstrap replicates of the variable of interest.
# S3 method for boot plot(x, index = 1, t0 = NULL, t = NULL, jack = FALSE, qdist = "norm", nclass = NULL, df, ...)
Arguments
| x | An object of class |
|---|---|
| index | The index of the variable of interest within the output of
|
| t0 | The original value of the statistic. This defaults to
|
| t | The bootstrap replicates of the statistic. Usually this will take
on its default value of |
| jack | A logical value indicating whether a jackknife-after-bootstrap plot is required. The default is not to produce such a plot. |
| qdist | The distribution against which the Q-Q plot should be drawn. At
present |
| nclass | An integer giving the number of classes to be used in the bootstrap
histogram. The default is the integer between 10 and 100 closest to
|
| df | If |
| ... | When |
Value
boot.out is returned invisibly.
Side Effects
All screens are closed and cleared and a number of plots are produced on the current graphics device. Screens are closed but not cleared at termination of this function.
Details
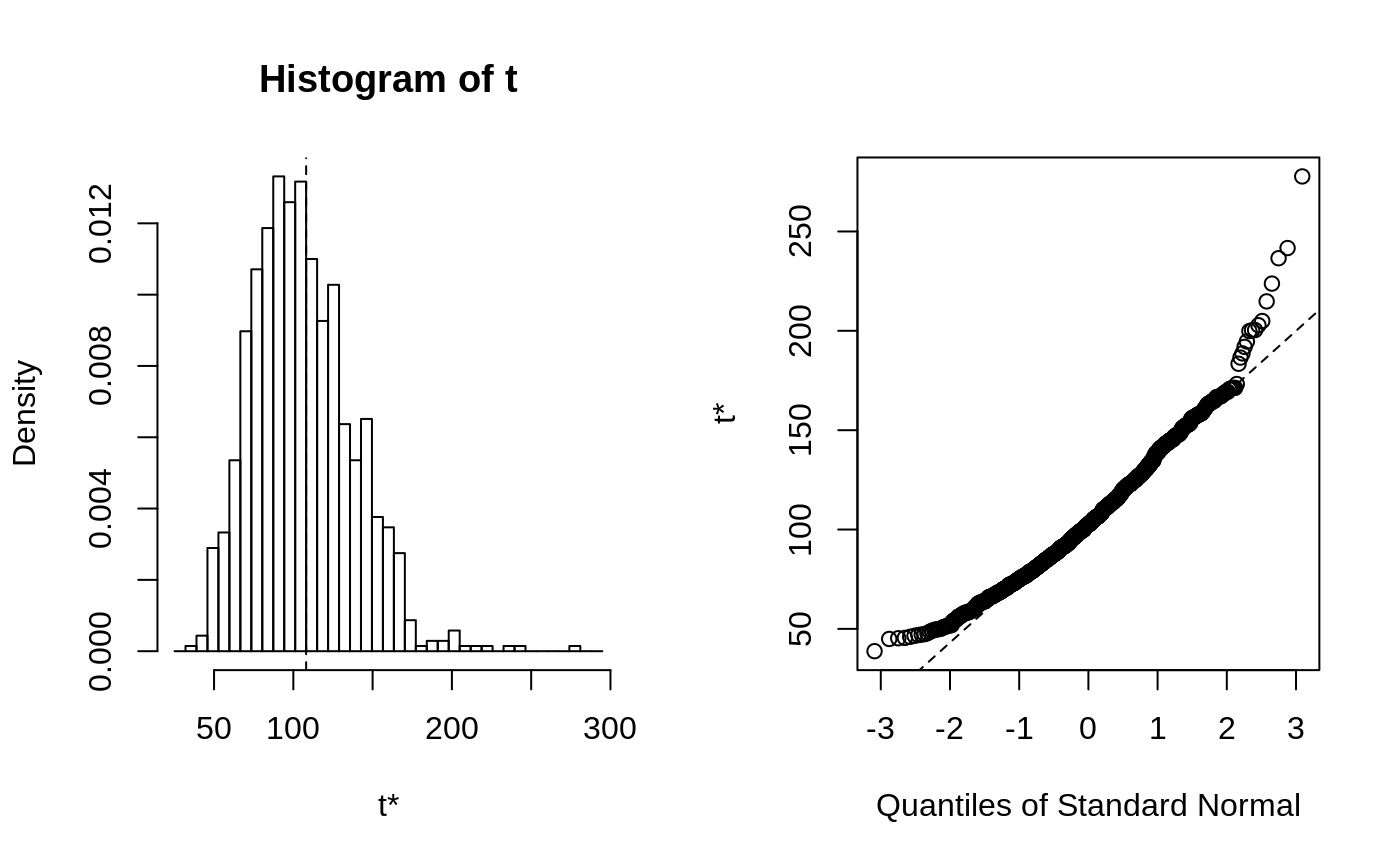
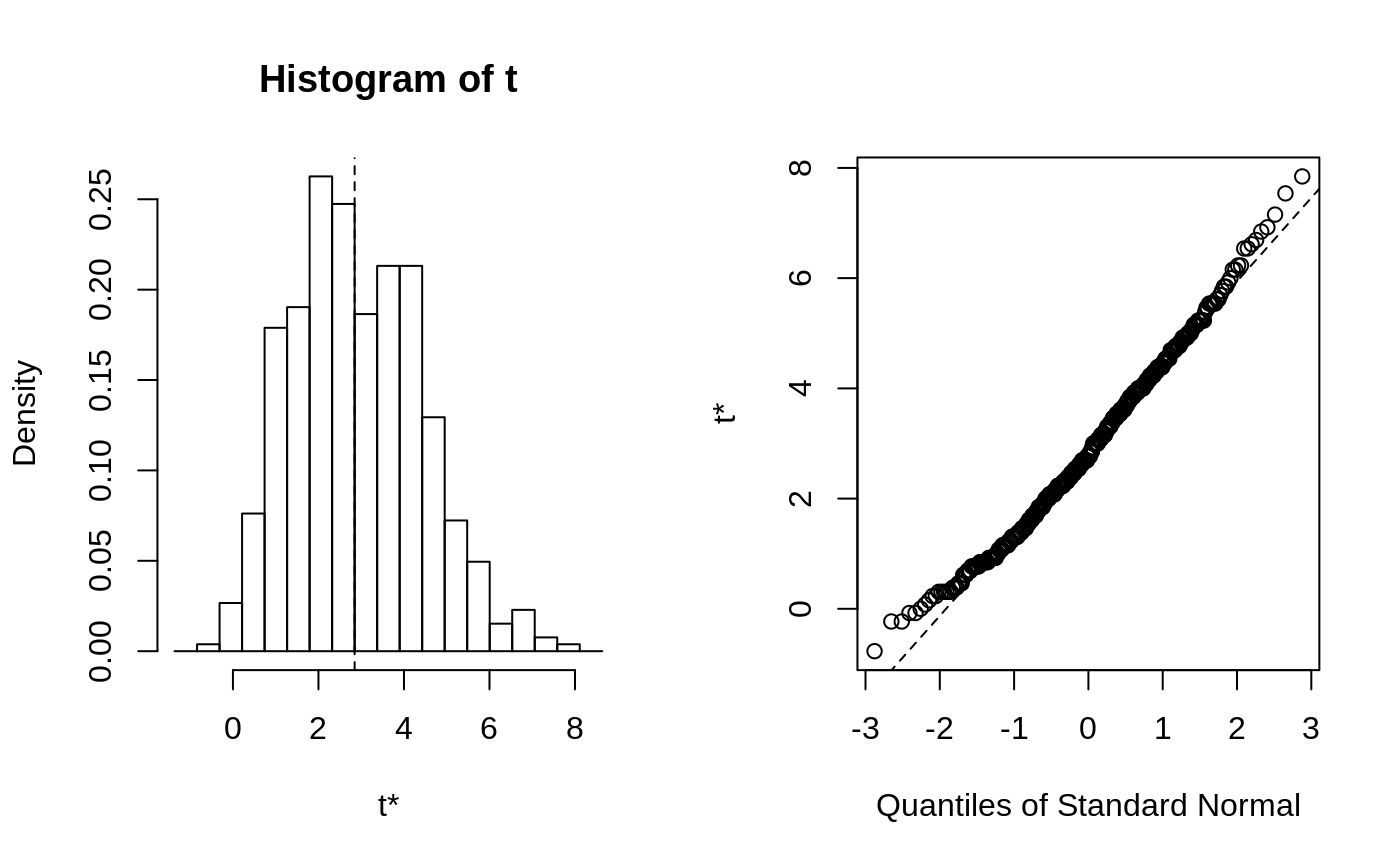
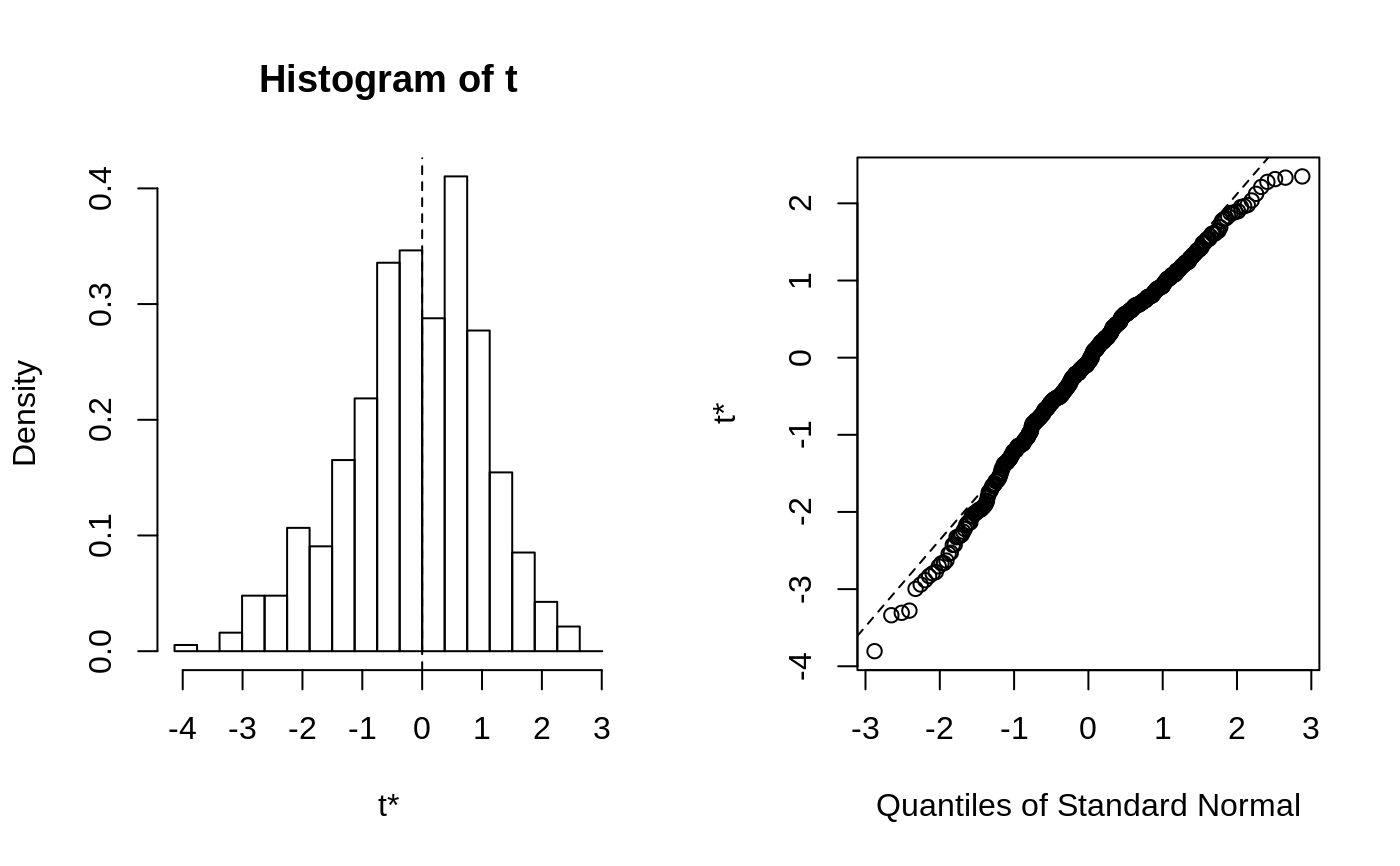
This function will generally produce two side-by-side plots. The left
plot will be a histogram of the bootstrap replicates. Usually the
breaks of the histogram will be chosen so that t0 is at a
breakpoint and all intervals are of equal length. A vertical dotted
line indicates the position of t0. This cannot be done if
t is supplied but t0 is not and so, in that case, the
breakpoints are computed by hist using the nclass
argument and no vertical line is drawn.
The second plot is a Q-Q plot of the bootstrap replicates. The order
statistics of the replicates can be plotted against normal or
chi-squared quantiles. In either case the expected line is also
plotted. For the normal, this will have intercept mean(t) and
slope sqrt(var(t)) while for the chi-squared it has intercept 0
and slope 1.
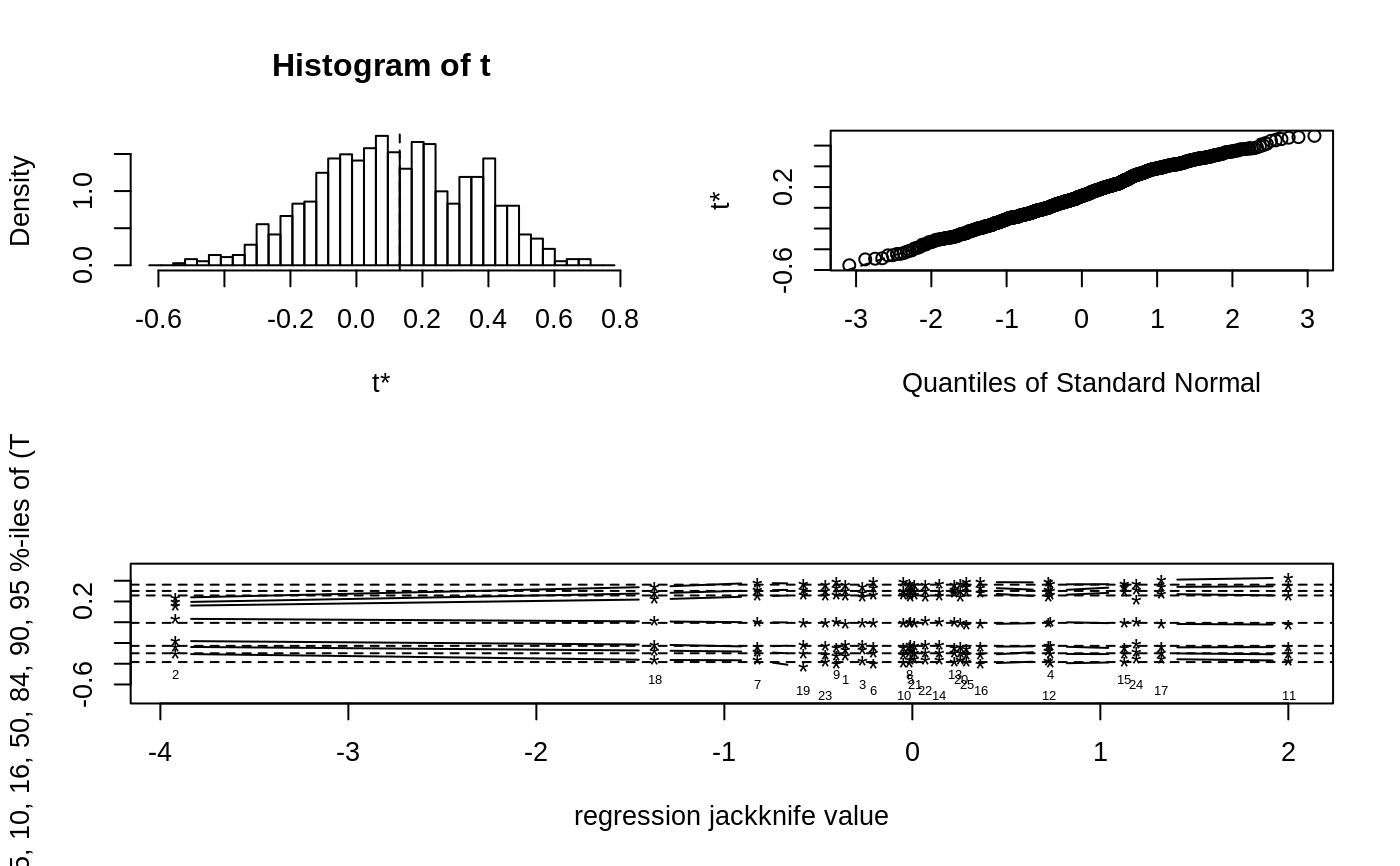
If jack is TRUE a third plot is produced beneath these
two. That plot is the jackknife-after-bootstrap plot. This plot may
only be requested when nonparametric simulation has been used. See
jack.after.boot for further details of this plot.
See also
Examples
# We fit an exponential model to the air-conditioning data and use # that for a parametric bootstrap. Then we look at plots of the # resampled means. air.rg <- function(data, mle) rexp(length(data), 1/mle) air.boot <- boot(aircondit$hours, mean, R = 999, sim = "parametric", ran.gen = air.rg, mle = mean(aircondit$hours)) plot(air.boot)# In the difference of means example for the last two series of the # gravity data grav1 <- gravity[as.numeric(gravity[, 2]) >= 7, ] grav.fun <- function(dat, w) { strata <- tapply(dat[, 2], as.numeric(dat[, 2])) d <- dat[, 1] ns <- tabulate(strata) w <- w/tapply(w, strata, sum)[strata] mns <- as.vector(tapply(d * w, strata, sum)) # drop names mn2 <- tapply(d * d * w, strata, sum) s2hat <- sum((mn2 - mns^2)/ns) c(mns[2] - mns[1], s2hat) } grav.boot <- boot(grav1, grav.fun, R = 499, stype = "w", strata = grav1[, 2]) plot(grav.boot)# now suppose we want to look at the studentized differences. grav.z <- (grav.boot$t[, 1]-grav.boot$t0[1])/sqrt(grav.boot$t[, 2]) plot(grav.boot, t = grav.z, t0 = 0)# In this example we look at the one of the partial correlations for the # head dimensions in the dataset frets. frets.fun <- function(data, i) { pcorr <- function(x) { # Function to find the correlations and partial correlations between # the four measurements. v <- cor(x) v.d <- diag(var(x)) iv <- solve(v) iv.d <- sqrt(diag(iv)) iv <- - diag(1/iv.d) %*% iv %*% diag(1/iv.d) q <- NULL n <- nrow(v) for (i in 1:(n-1)) q <- rbind( q, c(v[i, 1:i], iv[i,(i+1):n]) ) q <- rbind( q, v[n, ] ) diag(q) <- round(diag(q)) q } d <- data[i, ] v <- pcorr(d) c(v[1,], v[2,], v[3,], v[4,]) } frets.boot <- boot(log(as.matrix(frets)), frets.fun, R = 999) plot(frets.boot, index = 7, jack = TRUE, stinf = FALSE, useJ = FALSE)