Recursive Partitioning and Regression Trees
rpart.RdFit a rpart model
rpart(formula, data, weights, subset, na.action = na.rpart, method, model = FALSE, x = FALSE, y = TRUE, parms, control, cost, ...)
Arguments
| formula | a formula, with a response but no interaction
terms. If this a a data frame, that is taken as the model frame
(see |
|---|---|
| data | an optional data frame in which to interpret the variables named in the formula. |
| weights | optional case weights. |
| subset | optional expression saying that only a subset of the rows of the data should be used in the fit. |
| na.action | the default action deletes all observations for which
|
| method | one of Alternatively, |
| model | if logical: keep a copy of the model frame in the result?
If the input value for |
| x | keep a copy of the |
| y | keep a copy of the dependent variable in the result. If
missing and |
| parms | optional parameters for the splitting function. |
| control | a list of options that control details of the
|
| cost | a vector of non-negative costs, one for each variable in the model. Defaults to one for all variables. These are scalings to be applied when considering splits, so the improvement on splitting on a variable is divided by its cost in deciding which split to choose. |
| ... | arguments to |
Details
This differs from the tree function in S mainly in its handling
of surrogate variables. In most details it follows Breiman
et. al (1984) quite closely. R package tree provides a
re-implementation of tree.
Value
An object of class rpart. See rpart.object.
References
Breiman L., Friedman J. H., Olshen R. A., and Stone, C. J. (1984) Classification and Regression Trees. Wadsworth.
See also
Examples
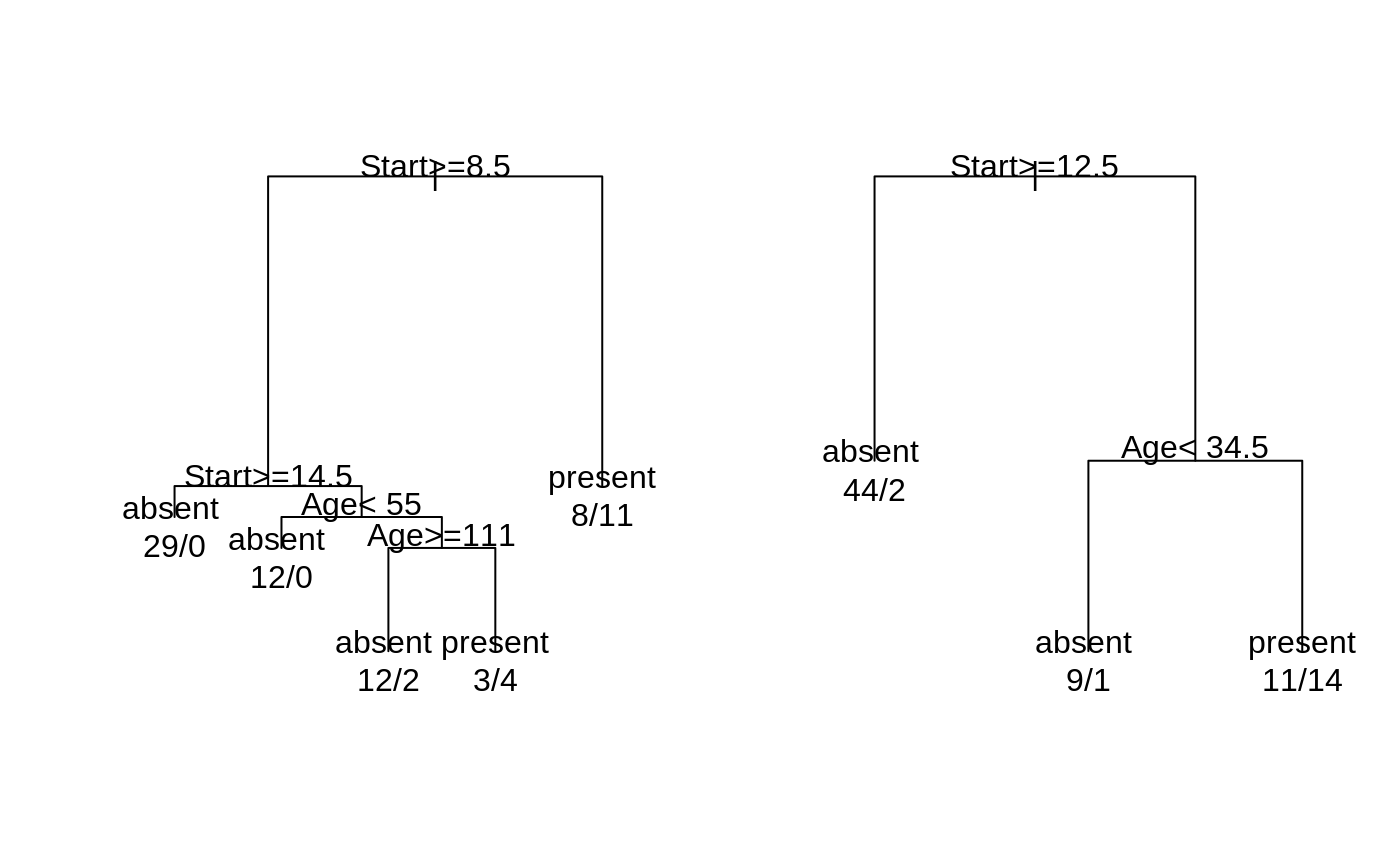
fit <- rpart(Kyphosis ~ Age + Number + Start, data = kyphosis) fit2 <- rpart(Kyphosis ~ Age + Number + Start, data = kyphosis, parms = list(prior = c(.65,.35), split = "information")) fit3 <- rpart(Kyphosis ~ Age + Number + Start, data = kyphosis, control = rpart.control(cp = 0.05)) par(mfrow = c(1,2), xpd = NA) # otherwise on some devices the text is clipped plot(fit) text(fit, use.n = TRUE) plot(fit2)